CNTNAP2 Heterozygous Missense Variants: Risk Factors for Autism Spectrum Disorder and/or Other Pathologies? - Giorgia Canali, Laurence Goutebroze, 2018
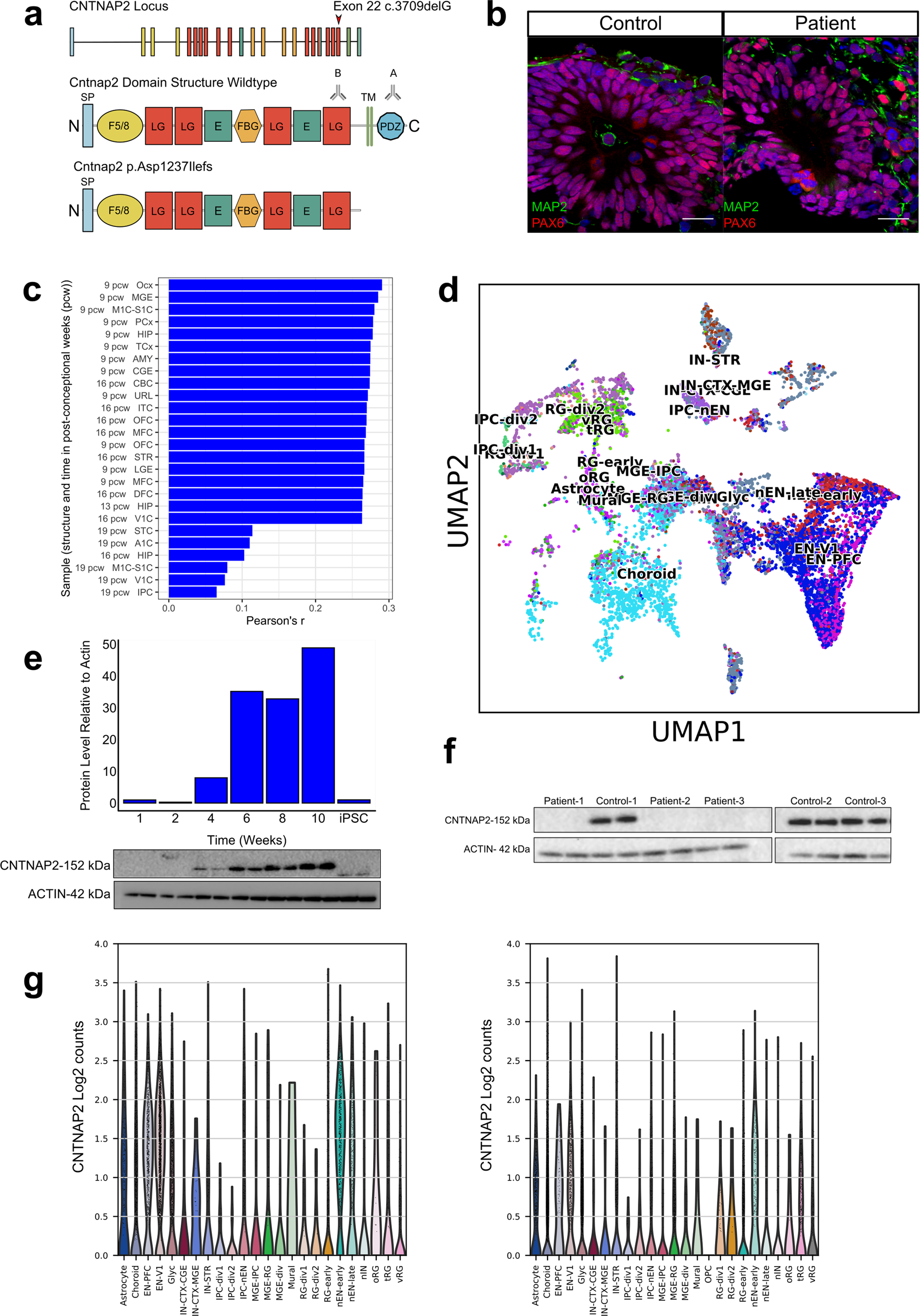
Cortical overgrowth in a preclinical forebrain organoid model of CNTNAP2-associated autism spectrum disorder | Nature Communications
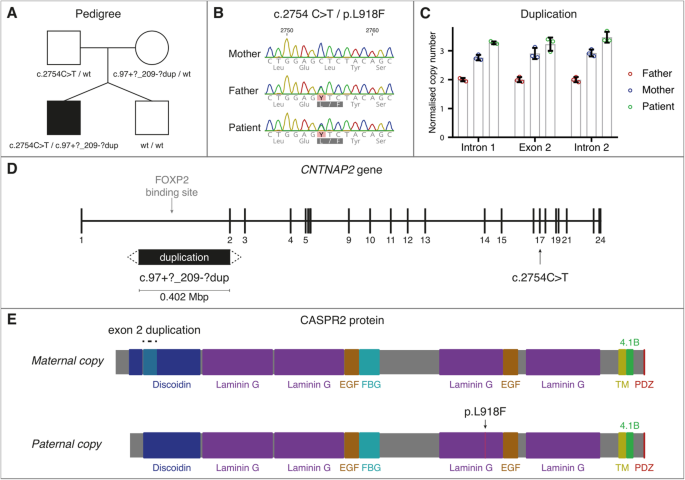
Hyperkinetic stereotyped movements in a boy with biallelic CNTNAP2 variants | Italian Journal of Pediatrics | Full Text

PDF) CNTNAP2 Heterozygous Missense Variants: Risk Factors for Autism Spectrum Disorder and/or Other Pathologies?

PDF) Differential impacts of Cntnap2 heterozygosity and Cntnap2 null homozygosity on axon and myelinated fiber development in mouse
No Evidence for Association of Autism with Rare Heterozygous Point Mutations in Contactin-Associated Protein-Like 2 (CNTNAP2), or in Other Contactin-Associated Proteins or Contactins | PLOS Genetics

Publications of the week – CNTNAP2, DEPDC5, and autism whole-genome sequencing | Beyond the Ion Channel

CNTNAP2 Heterozygous Missense Variants: Risk Factors for Autism Spectrum Disorder and/or Other Pathologies? - Giorgia Canali, Laurence Goutebroze, 2018

Hippocampal gamma and sharp-wave ripple oscillations are altered in a Cntnap2 mouse model of autism spectrum disorder - ScienceDirect
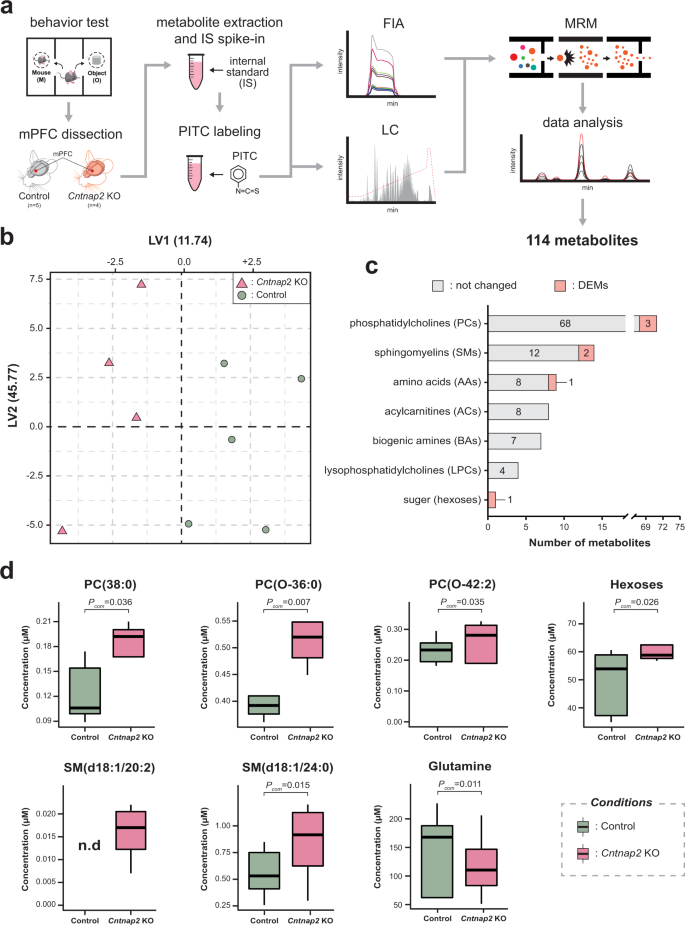
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry
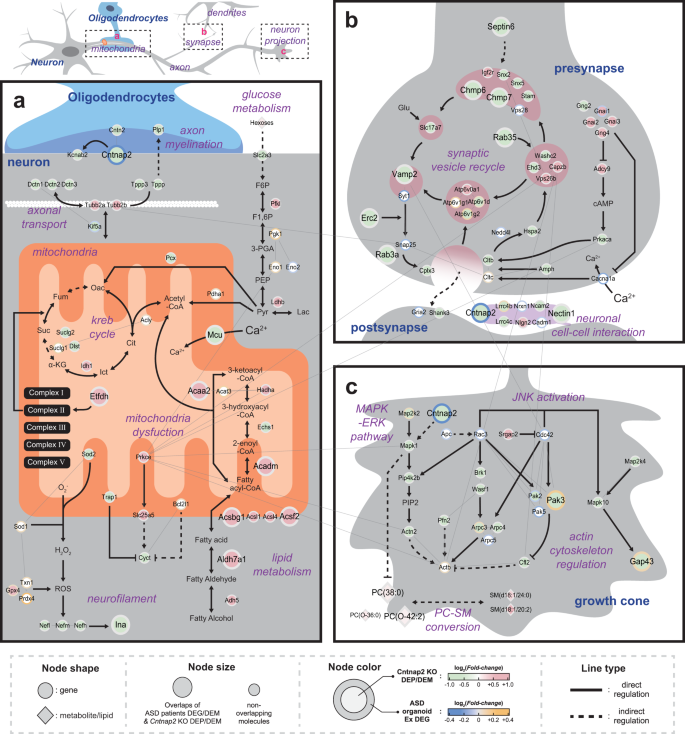
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry

Frontiers | Dysregulation of Parvalbumin Expression in the Cntnap2−/− Mouse Model of Autism Spectrum Disorder
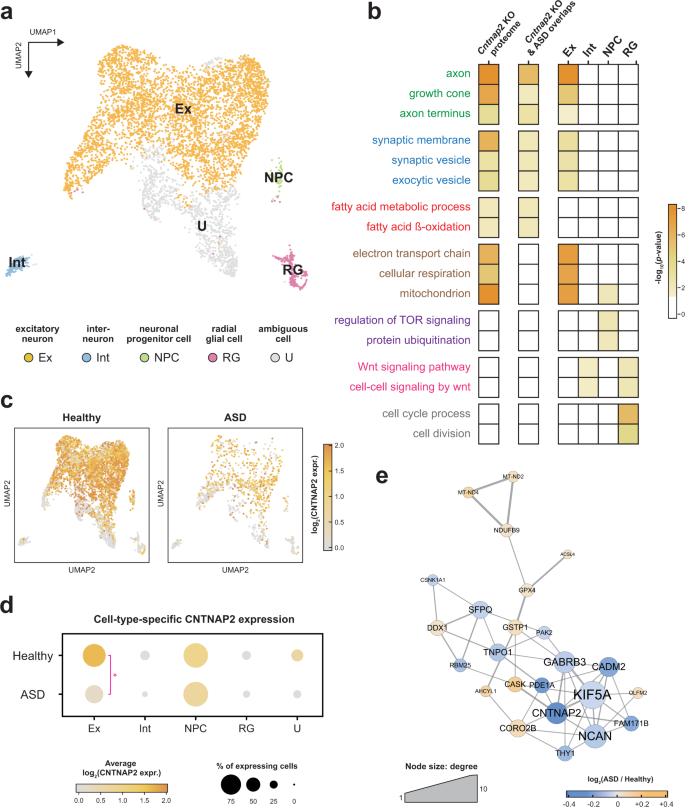
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry
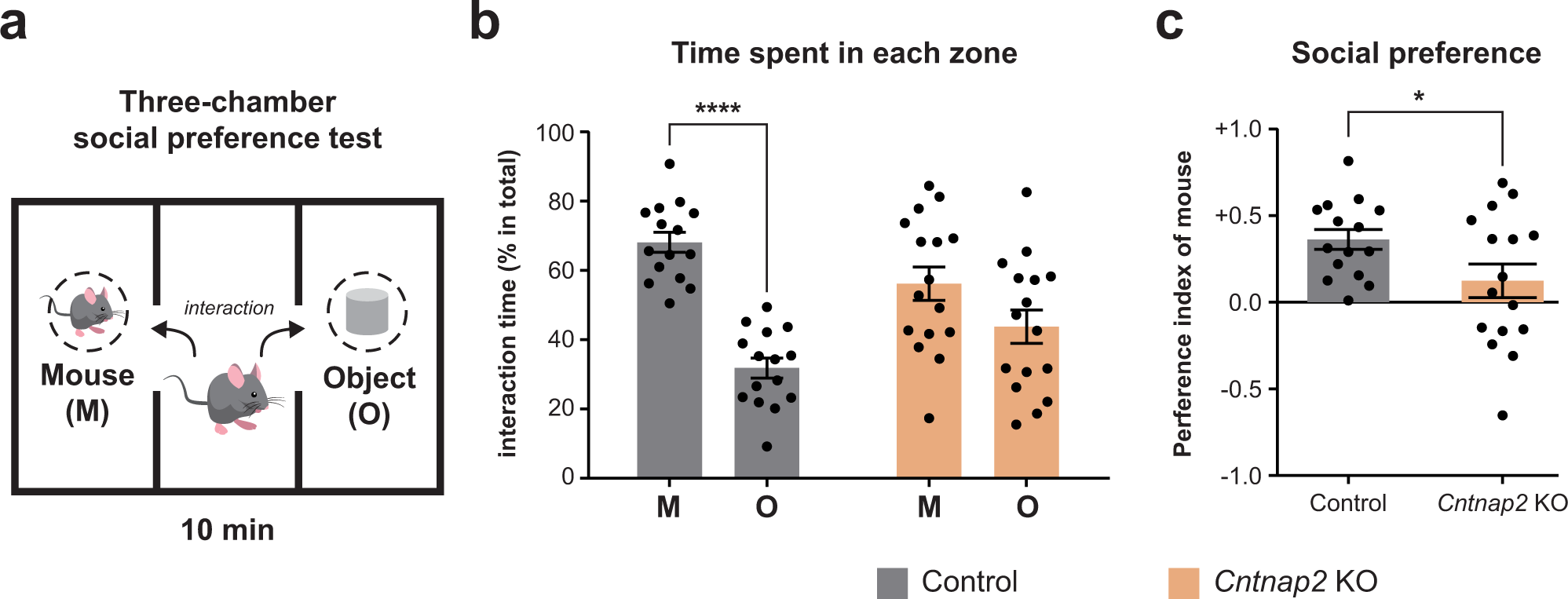
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry
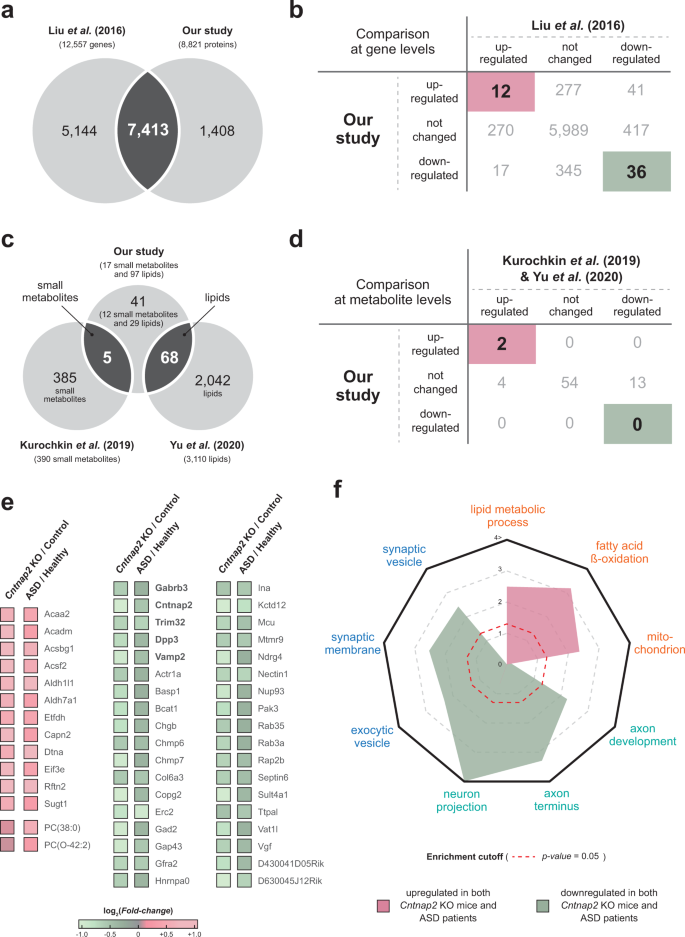
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry

Absence of CNTNAP2 Leads to Epilepsy, Neuronal Migration Abnormalities, and Core Autism-Related Deficits: Cell

JCM | Free Full-Text | Altered Blood Brain Barrier Permeability and Oxidative Stress in Cntnap2 Knockout Rat Model
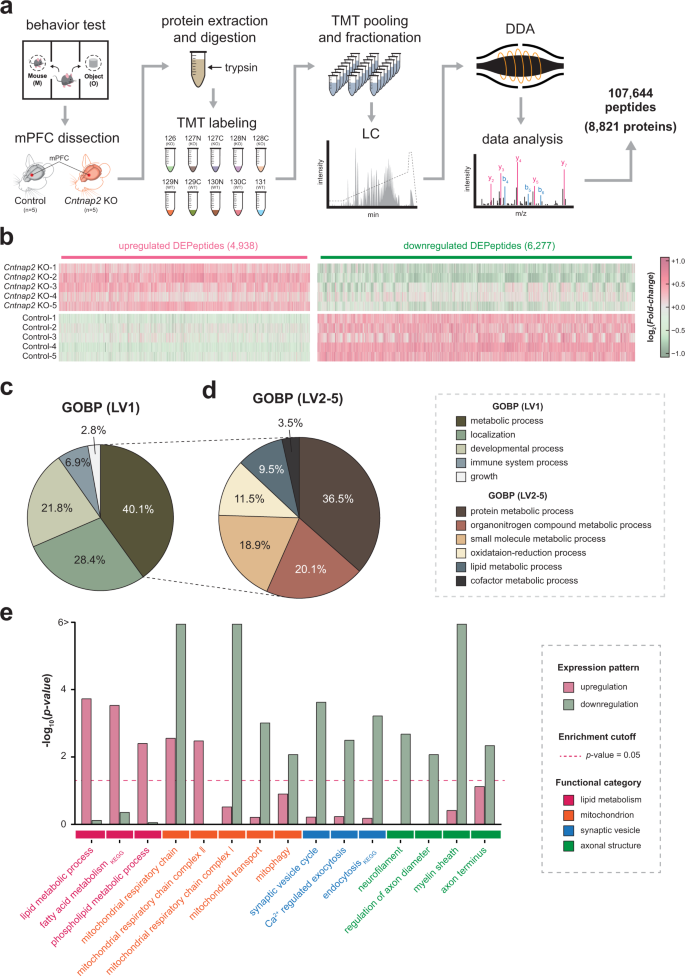
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry
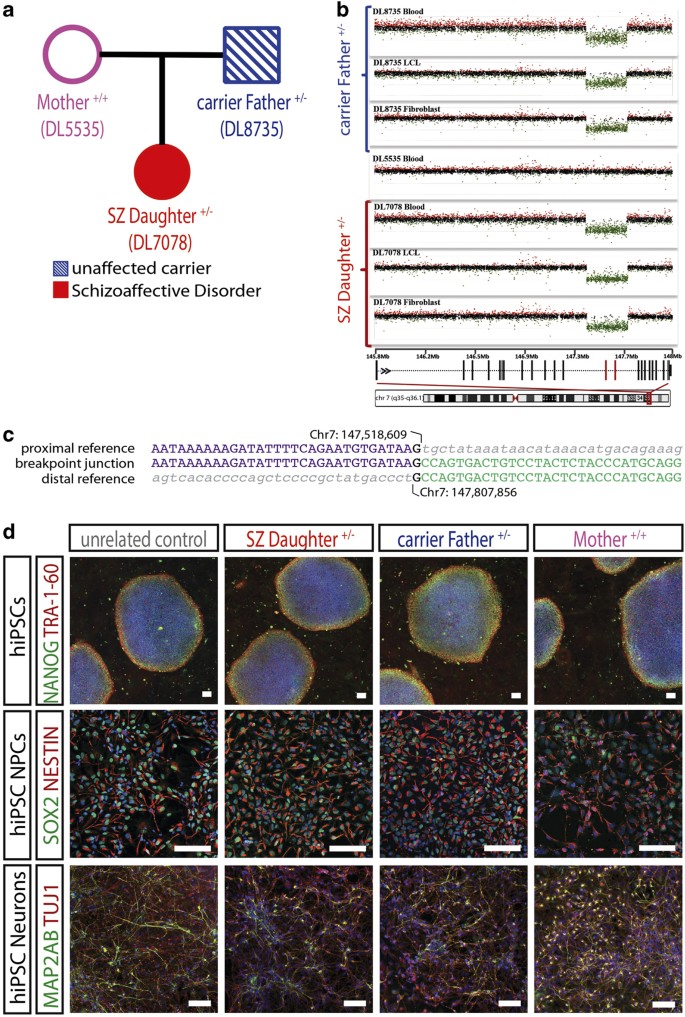
Characterization of molecular and cellular phenotypes associated with a heterozygous CNTNAP2 deletion using patient-derived hiPSC neural cells | Schizophrenia